iNDI-PD
ASAP is supporting iNDI-PD (an expansion to the original iPSC Neurodegenerative Disease Initiative (iNDI) and anchored within the NIH Center for Alzheimer’s and Related Dementias (CARD) to make iPSC lines that are relevant to different neurodegenerative diseases, including Parkinson’s disease. Lines generated through iNDI-PD are accessible to all researchers interested in understanding Parkinson’s disease to discover novel insights into the molecular biology of disease and to ensure that additional mutations relevant to Parkinson’s research are also included in their efforts.
Induced Pluripotent Stem Cells
Induced pluripotent stem cells (iPSCs) are derived from adult cells such as skin or blood and reprogrammed back into pluripotent stem cells. These appealing research tools create the ability to study a disorder in disease-relevant cell types from real patients. However, donors who provide these cells carry innumerable genetic differences, making it hard to pinpoint whether an observation is due to a disease-causing mutation or something else in the patient’s genetic makeup. iNDI introduces disease-causing mutations to a set of cell lines that all have the same genetic background (called isogenic lines). By incorporating Parkinson’s disease data in the neurodegenerative resource bank, we are also enabling researchers to draw connections across diagnostic lines.
Overview of iNDI-PD
iNDI is a large iPSC genome-engineering initiative, with more than 600 engineered daughter lines to be generated from each parental line. iPSC lines are CRISPR/Cas9 engineered to express one or two copies of single nucleotide variants (SNVs) that are relevant to these diseases in a common cellular background. Additional control lines are also developed for each SNV line, in which the disease-relevant mutation is reverted back to the wild-type allele. Companion lines will include revertants, Halo-tagged knockins, and knockouts. iNDI-PD will develop Parkinson’s disease-relevant iPSC lines across three phases.

Phase 1
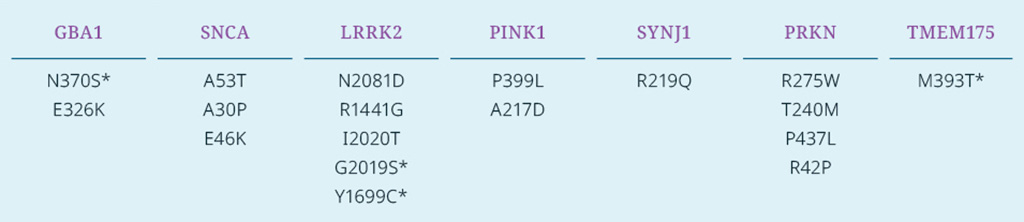
iNDI-PD focused on SNVs in well-established Parkinson’s disease-relevant genes like SNCA, LRRK2, GBA1, PINK1, and PRKN. Over 15 trio sets (of homozygous SNV, heterozygous SNV, and revertant lines) have been generated, quality-controlled, and made available to the research community through The Jackson Laboratory (JAX).
Phase 2
iNDI-PD will generate gene knockout lines and lines harboring genetic variants recently reported in the Parkinson’s disease/Dementia with Lewy Bodies literature.
Phase 3 (Coming Soon)
iNDI-PD will expand the number of SNV and knockout lines, based on recent findings and nominations from researchers in the ASAP community (Collaborative Research Network, Global Parkinson’s Genetics Program, and Parkinson’s Progression Markers Initiative).

iNDI-PD Outputs in the ASAP Catalog
Our initiative is accelerating the pace of discovery for Parkinson’s disease through collaboration, research-enabling resources, and data sharing. Check out our Catalog to view all ASAP-supported lab materials from iNDI-PD.

